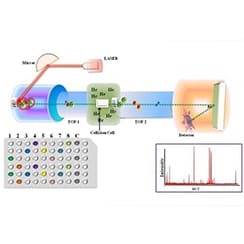
Mass Spectroscopy
At least a 100-year-old technique, Mass spectroscopy or Mass spectrometry has an important application in Proteomics. This technique involves the use of a machine which ionizes the sample and then separates or sorts them based on their mass-to-charge ratio. The substance to be analyzed is known as an analyte and this could be gas, solid or liquid.
The first step referred as 'Ionization' is the bombarding of the sample particles with the electrons. This leads to the acquisition of a positive charge as electrons from the outermost shell of the sample molecule are knocked out. This Ionization can be brought about in various ways. For example, the Electron Ionization (EI), Chemical Ionization (CI), the Fast Atom Bombardment (FAB), Electrospray ionization (ESI) etc. The second stage is termed as Mass filtration. Here the mass analyzer separates the ions based on the charge to mass ratio. 'Detection' is the next stage. This stage can apply the use of various types of detectors. The most commonly used are electron multipliers which have dynodes bars that are made of a material that releases electrons when impacted by particles. A plot known as the mass spectrum will be generated with intensity (the current generated via electron multiplier) on the Y-axis and the mass/charge ratio on the X.
Mass spectroscopy finds its application in protein characterization pharmacokinetics, isotope dating and in separation techniques like Chromatography.
No Products


